Ultimate Resource On Crispr And It’s Various Applications For Advanced Gene-Editiing
As new gene editing tools raise the prospect of engineering desired human traits, researchers are determined to educate the public. Ultimate Resource On Crispr And It’s Various Applications For Advanced Gene-Editiing
In the early decades of the 20th century, prominent American scientists and physicians were involved in the eugenics movement, promoting reproduction among those seen as being more genetically fit. They also helped to lobby state legislatures to pass laws compelling the involuntary sterilization of people deemed genetically inferior.
In the 1927 case of Buck v. Bell, the U.S. Supreme Court upheld the constitutionality of Virginia’s program. As Justice Oliver Wendell Holmes, Jr. , notoriously declared in the decision, “Three generations of imbeciles is enough.”
In time, however, eugenics lost credibility and support, having become associated with atrocities committed by the Nazis. Many came to see the idea that humans might change their DNA to control the genetic future as both scientifically unlikely and immoral.
Related:
Bitcoin Community Leaders Join Longevity Movement
Bitcoiners Are Now Into Fasting. Read This Article To Find Out Why
The Definitive Guide On Fasting, Autophagy, And Caloric Restriction
Fasting 101 Rebooting The Body’s Hard Drive Rejuvenating, Life-Extending And Removes Deadly Toxins
What Is Dopamine Fasting? Meet The Dangerous Fad Among Silicon Valley’s Tech Geniuses
The invention of Crispr has now brought that power into the realm of the possible, and from the start, excitement about its potential to do good has been tempered by fear about the possibility of causing irrevocable harm. “Geneticists have a historical burden,” says Mr. Greely. “Their science was used in ways that turned out to be deeply unscientific.” He does not think scientists should be blamed for such misuse, “but they should be held responsible for giving some thought to how this might go wrong, and talking about how to maximize benefits and minimize risks.”
Jennifer Doudna, a professor at the University of California, Berkeley, is one of the inventors of the Crispr tool. She has recounted a nightmare she had about the technology. In the dream, a colleague told her that somebody wanted to talk to her about gene editing. When she entered the room, the person waiting to meet her was Adolf Hitler. Dr. Doudna and her colleagues hoped Crispr might ultimately save lives, she wrote. But the nightmare was a reminder of “all of the ways in which our hard work might be perverted.”
Daniel Kevles, a professor of history emeritus at Yale University and the author of a history of eugenics, says that in current discussions about Crispr, people recoil from any association with eugenics, because the term is so closely linked to state-sponsored abuses. In the U.S., involuntary programs like the one in Buck v. Bell continued through the 1970s, sterilizing at least 60,000 people. But in the Crispr era, Prof. Kevles says, the moral dilemmas surrounding genetic manipulation are less likely to be caused by government initiatives than by “private eugenics”: consumers who want to use Crispr or other genetic technologies to give their children advantages in life.
Despite advances in genetic technology, scientists still do not have a reliable way to “manipulate human heredity to guarantee the birth of a child that can put a basketball through a hoop at 30 feet or perform in Carnegie Hall,” says Prof. Kevles. “If and when that happens, people will want to make use of it. We live in a consumer culture, and people want the best for their kids.”
Many of the institutions involved in research on genetic engineering are already doing outreach to the public, explaining how gene editing works. But now they are trying to do more on the ethical front, too. In one high-profile effort, Harvard Medical School’s Personal Genetics Education Project has initiated a Genetics Consortium, whose activities will include offering education programs across the country, from rural high schools to urban churches.
Ting Wu, a professor of genetics at Harvard Medical School and one of the leaders of the Genetics Consortium, says that for the pilot effort, they focused on seven institutions that have played important roles in the current genetic revolution. As it turned out, some of them have historical connections to the eugenics movement. A 1914 article in the Journal of Heredity, for example, included a list of colleges and universities that offered courses on eugenics, and Consortium members such as Harvard, the University of California and the University of Washington were on the list.
Michael Snyder, chairman of the department of genetics at Stanford University School of Medicine and a member of the Consortium, says that its initial focus is going to be on educating people about what genetic technology can and cannot do. Using Crispr to change complicated traits is still not imminent, he said. This month, three reports were published by scientists arguing over whether researchers successfully used Crispr to edit out a single gene mutation in human embryos linked to a deadly heart disease. To significantly manipulate someone’s intelligence or athletic ability, Dr. Snyder says, likely would require thousands of gene changes. And even then, genetics remains only part of the story. Diet, environment and training also play a role.
Still, Dr. Snyder says, these are questions that the Consortium will need to address. If it becomes possible, he could imagine parents wanting to tweak the genome of an embryo with, say, a growth hormone deficiency, in order to make the future child taller. But who gets to decide what sort of height deficiency justifies intervention? What, he asks, if “you are not impaired in any way other than you are not going to be the center of the basketball team?”
At Harvard’s Personal Genetics Education Project, the Buck v. Bell sterilization case is discussed in a lesson plan designed for use in classrooms and teacher training workshops. The case not only highlights how science can be misused but the conviction of many proponents of eugenics that “they were doing good,” says Dr. Wu. “Do I think there are people who are going to do things that 100 years from now we will be shocked they did but today they believe it is the right thing to do?” she asks. “Sure, I think that’s a possibility. The more eyes we have on this, the more arguments about this, the better we will all be.”
Crispr Used To Repair Gene Mutation in Dogs With Muscular Dystrophy
Gene-editing tool holds potential to edit DNA of patients with deadly disease.
Researchers used a gene-editing tool to repair a gene mutation in dogs with Duchenne muscular dystrophy, an important step in efforts to someday use the tool to edit DNA in people with the same fatal disease.
In a study published Thursday in the journal Science, researchers at UT Southwestern Medical Center in Dallas and the Royal Veterinary College in London reported that they used the Crispr gene-editing system in four dogs to restore production of dystrophin, a protein crucial for healthy muscle function.
Duchenne is caused by gene mutations that prevent production of dystrophin. Without enough dystrophin, patients’ muscles deteriorate. Patients typically lose the ability to walk by their early teens and die by their 20s and 30s. About 20,000 children, mostly boys, are diagnosed around the world every year with Duchenne.
Scientists have tried many approaches to stop the disease. In 2016, the U.S. Food and Drug Administration approved the first drug, Sarepta Therapeutics Inc.’s Exondys 51, a weekly injection that treats a particular form of Duchenne.
More recently, small numbers of people have been dosed with experimental gene therapies that attempt to bolster dystrophin production by delivering a form of the protein to cells. There are over 45 companies pursuing strategies to treat Duchenne or its symptoms, says Pat Furlong, CEO of the nonprofit Parent Project Muscular Dystrophy, which supported some of UT Southwestern’s early Crispr work in mice.
What makes the Crispr gene-editing approach in the dog study so alluring to many in the field is the prospect of tackling the underlying genetic cause of Duchenne. “Crispr is the next step,” says Ms. Furlong, “because it can permanently change people’s DNA.”
Crispr, which stands for clustered regularly interspaced short palindromic repeats, serves as the immune system of bacteria. Scientists adapted Crispr and the Cas9 enzyme it produces to cut DNA at specific points, allowing a repair or insertion of a new genetic sequence. Its development has raised hopes for curing genetic diseases, but has also sparked concerns about unintentional cuts and deletions of DNA.
Stanley F. Nelson, father of a boy with Duchenne and professor and co-director of the Center for Duchenne Muscular Dystrophy at UCLA, said much more data is needed before moving from dogs to having a clinical trial with people.
“You can’t be certain at this point that in trying to repair billions and billions of muscle cells, you won’t induce other catastrophic problems,” he said. Dr. Nelson also consults for Solid Biosciences Inc., a company pursuing gene therapy for Duchenne. He wasn’t involved in the dog study.
In the Science study, scientists programmed the Crispr system to cut the dogs’ DNA at a precise spot on the dystrophin gene. The cells repaired the cut, enabling dystrophin production to be restored to levels that ranged from 3% to 90% of normal, depending on the muscle. Most importantly, the gene edit restored dystrophin production in both the heart and diaphragm, essential for breathing.
“It’s like putting a good spare tire on a car. It’s not as good as the original, but it gets you where you want to go,” said Eric Olson, director of UT Southwestern’s Hamon Center for Regenerative Science and Medicine and senior author of the paper.
Dr. Olson, who is also founder and chief scientific adviser of Exonics Therapeutics Inc., which licensed the technology from UT Southwestern and helped fund the dog studies, said next steps involve testing Crispr in more dogs and observing them for a year or more. If the approach works in the dogs, he said researchers hope to try Crispr in a clinical trial with people with Duchenne.
Related Article: Meet The Scientists Bringing Extinct Species Back From The Dead (#GotBitcoin?)
Updated: 4-11-2022
AI-Designed Protein Can Awaken Silenced Genes One-By-One
Gene therapy technique designed by the University of Washington can toggle on individual genes that regulate cell growth, development and function.
By combining CRISPR technology with a protein designed by AI, researchers from the University of Washington School of Medicine in Seattle have been able to awaken individual dormant genes by disabling the chemical “off switches” that silence them through gene therapy.
Longevity.Technology: This gene therapy approach, which is described in Cell Reports, will allow researchers to understand the role individual genes play in normal cell growth and development and diseases such as cancer; it will also broaden our understanding of aging at a genetic level. The new technique controls gene activity without altering the DNA sequence of the genome by targeting the epigenome – the chemical tags and modifications that help package genes in our chromosomes and regulate their activity. Epigenetic modifications affect gene activity and we collect them over time – this build-up of epigenetic markers contributes to aging, but can also affect of the health of future generations as they can be inherited.
Targeting the epigenome was an attractive prospect for the Washington team. “The beauty of this approach is we can safely upregulate specific genes to affect cell activity without permanently changing the genome and cause unintended mistakes,” Shiri Levy, a postdoctoral fellow in UW Institute for Stem Cell and Regenerative Medicine (ISCRM) and the lead author of the paper, says [1].
In the gene therapy research, Levy and her colleagues focused on a complex of proteins called PRC2 that silences genes by attaching a small molecule, called a methyl group, to a protein that packages genes called histones.
PRC2 is active throughout development but plays a particularly important role during the first days of life when embryonic cells differentiate into the various cell types that will form the tissues and organs of the growing embryo.
PRC2 can be blocked with chemicals, but they are imprecise, affecting PRC2 function throughout the genome. The goal of the UW researchers was to find a way to block PRC2 so that only one gene at a time would be affected.
To do this, David Baker and his colleagues used AI to create a protein that would bind to PRC2 and block a protein the PRC2 uses to modify the histones. Ruohola-Baker and Levy then fused this designed protein with a disabled version of a protein called Cas9.
Cas9 is the protein used in the gene editing process called CRISPR – Cas9 binds and uses RNA as an address-tag. The system allows scientists, by synthesising a specific “address-tag” RNA, to bring Cas9 to a precise location in the genome and cut and splice genes at very specific sites.
However, in this experiment the researchers disabled the cutting function of the Cas9 protein – calling it dCas9, for “dead”.
This meant the genomic DNA sequence remained unaltered. However, it can no longer cut, dCas9 can still deliver, functioning as a transport for cargo and delivering it to a specific location. The AI-designed blocking protein was the cargo of the dCas9-RNA construct.
“DCas9 is like UBER,” explains Levy, “It will take you anywhere on the genome you want to go. The guide RNA is like a passenger, telling the UBER where to go [1].”
In the new paper, Levy and her colleagues show that by using this technique, they were able to block PRC2 and selectively turn on four different genes. They were also able to show they could transdifferentiate induced pluripotent stem cells to placental progenitor cells by simply turning on two genes [2].
“This technique allows us to avoid bombarding cells with various growth factors and gene activators and repressors to get them to differentiate,” Levy explains. “Instead, we can target specific sites on the gene transcription promoters’ region, lift those marks and let the cell do the rest in an organic, holistic manner [1].”
Finally, the researchers were able to show how the technique can be used to find the location of specific PRC2-controlled regulatory regions from where individual genes are activated – the location of many of these are unknown. In this case, they identified a promoter region –called a TATA box – for a gene called TBX18.
Although current thinking is that these promotor regions are close to the gene, within in 30 DNA base pairs, they found for this gene the promoter region was more than 500 base pairs away.
“This was a very important finding,” said Ruohola-Baker. “TATA boxes are scattered throughout the genome, and current thinking in biology is that the important TATA boxes are very close to the gene transcription site and the others don’t seem to matter. The power of this tool is that it can find the critical PRC2 dependent elements, in this case TATA boxes that matter [1].”
Epigenetic modifications decorate broad regions of the genome in normal and abnormal cells. However, the minimal functional unit for the epigenetic modification remains poorly understood, Ruohola-Baker notes:
“With these two advances, AI-designed proteins and CRISPR technology, we can now find the precise epigenetic marks that are important for gene expression, learn the rules and utilize them to control cell function, drive cell differentiation and develop 21st century therapies [1].”
[1] https://bit.ly/3usm1x2
[2] https://www.cell.com/cell-reports/fulltext/S2211-1247(22)00184-X
Updated: 12-11-2022
Base Editing: Revolutionary Therapy Clears Girl’s Incurable Cancer
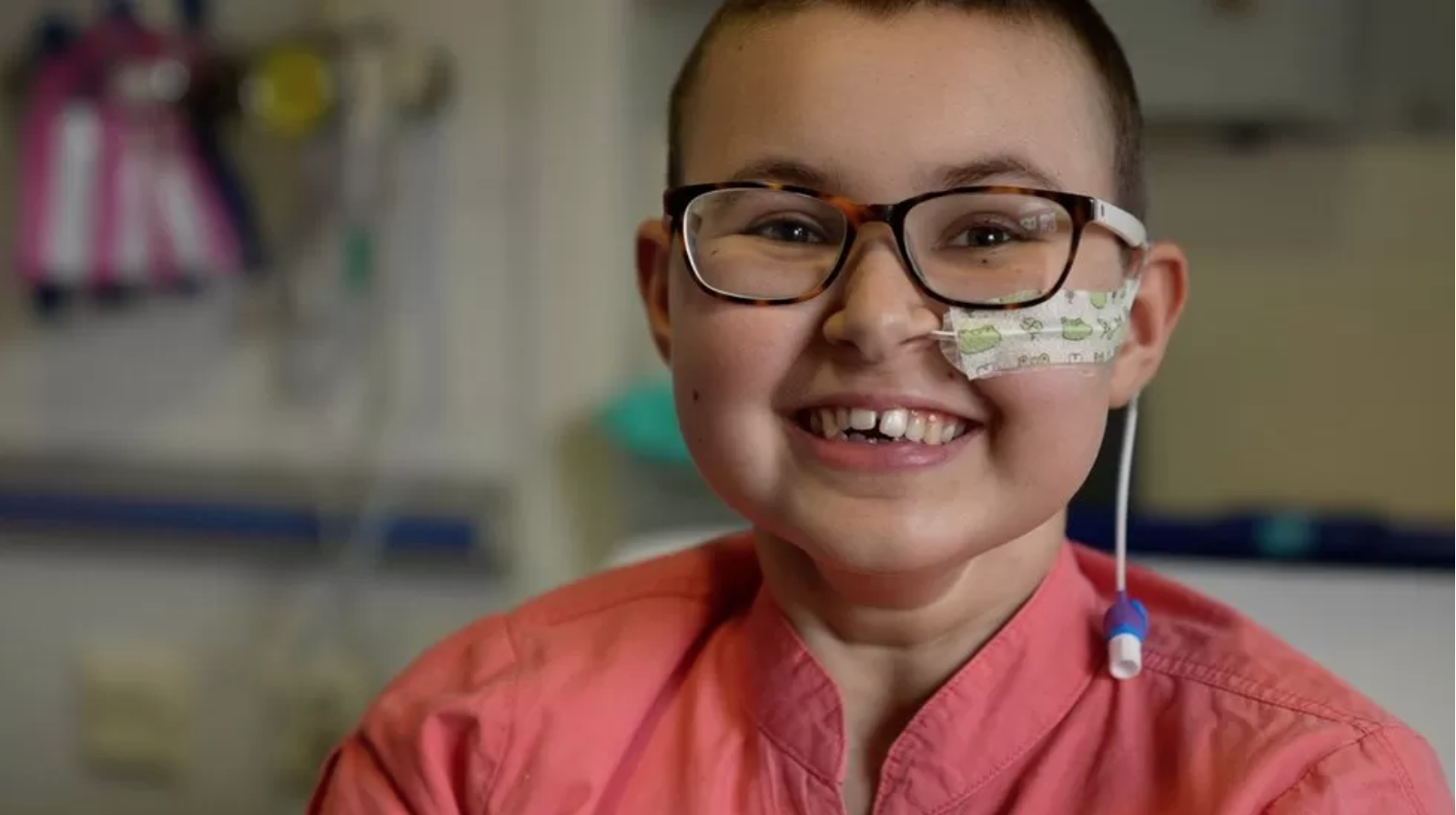
A teenage girl’s incurable cancer has been cleared from her body in the first use of a revolutionary new type of medicine.
All other treatments for Alyssa’s leukaemia had failed.
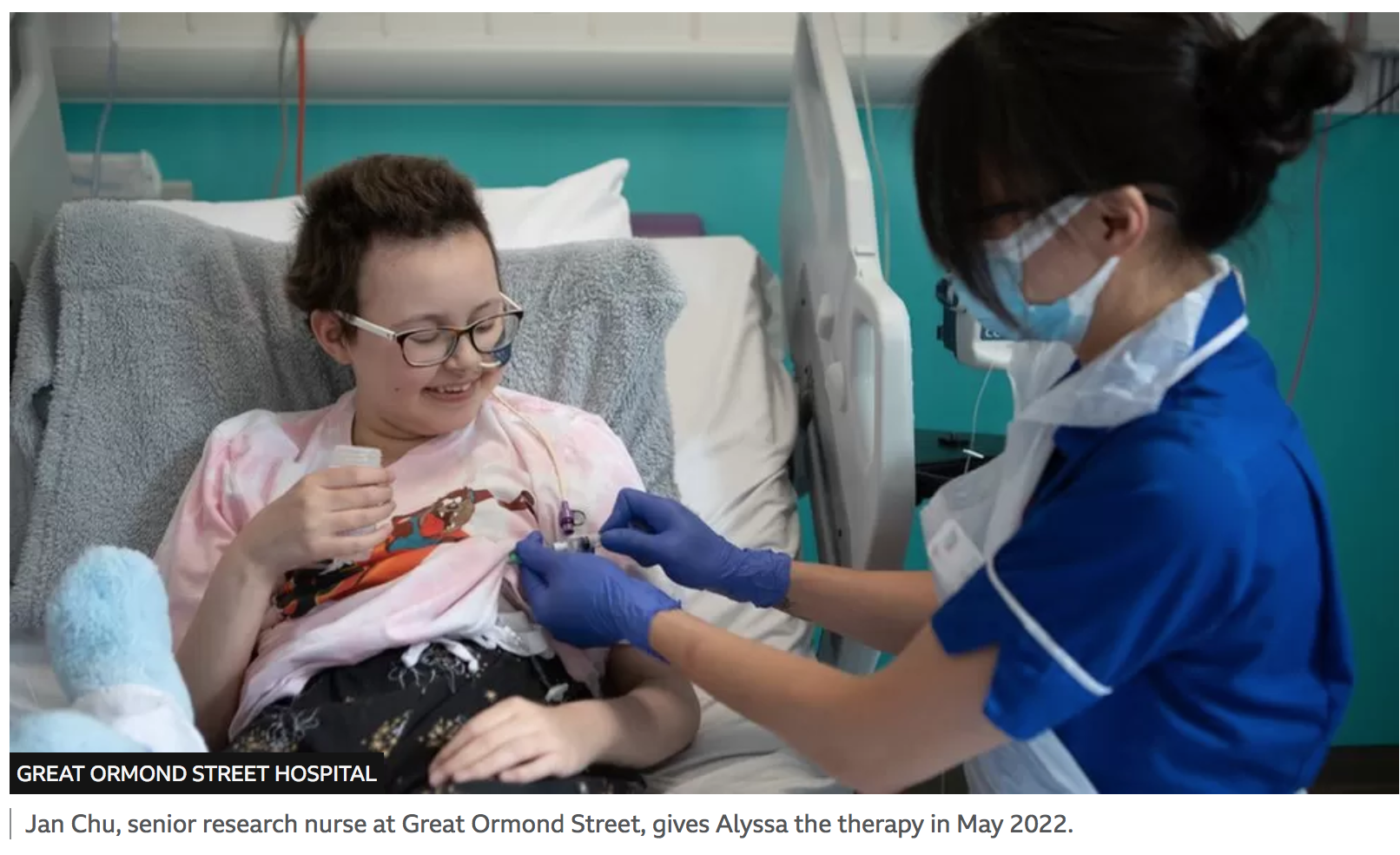
So doctors at Great Ormond Street Hospital used “base editing” to perform a feat of biological engineering to build her a new living drug.
Six months later the cancer is undetectable, but Alyssa is still being monitored in case it comes back.
Alyssa, who is 13 and from Leicester, was diagnosed with T-cell acute lymphoblastic leukaemia in May last year.
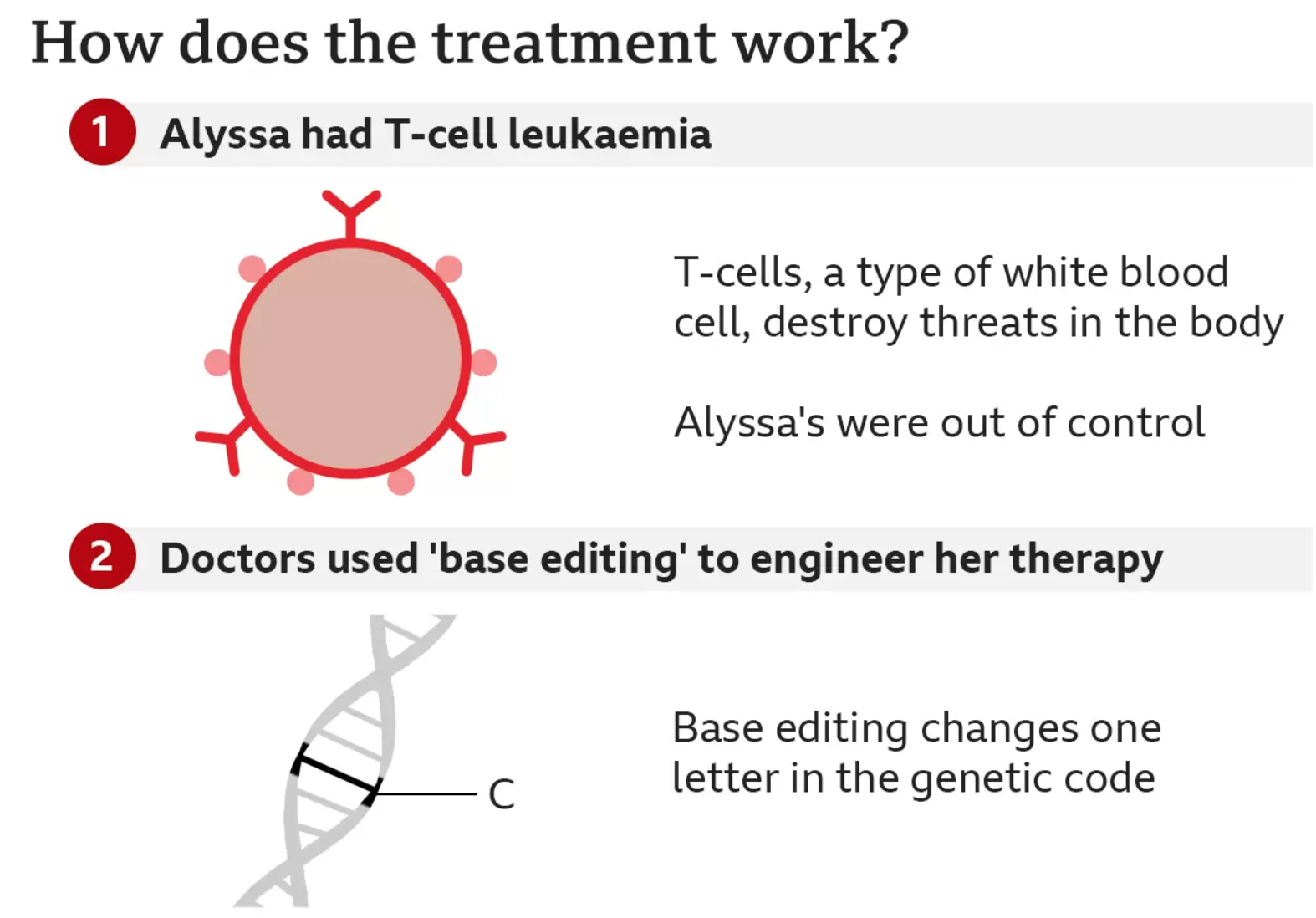
T-cells are supposed to be the body’s guardians – seeking out and destroying threats – but for Alyssa they had become the danger and were growing out of control.
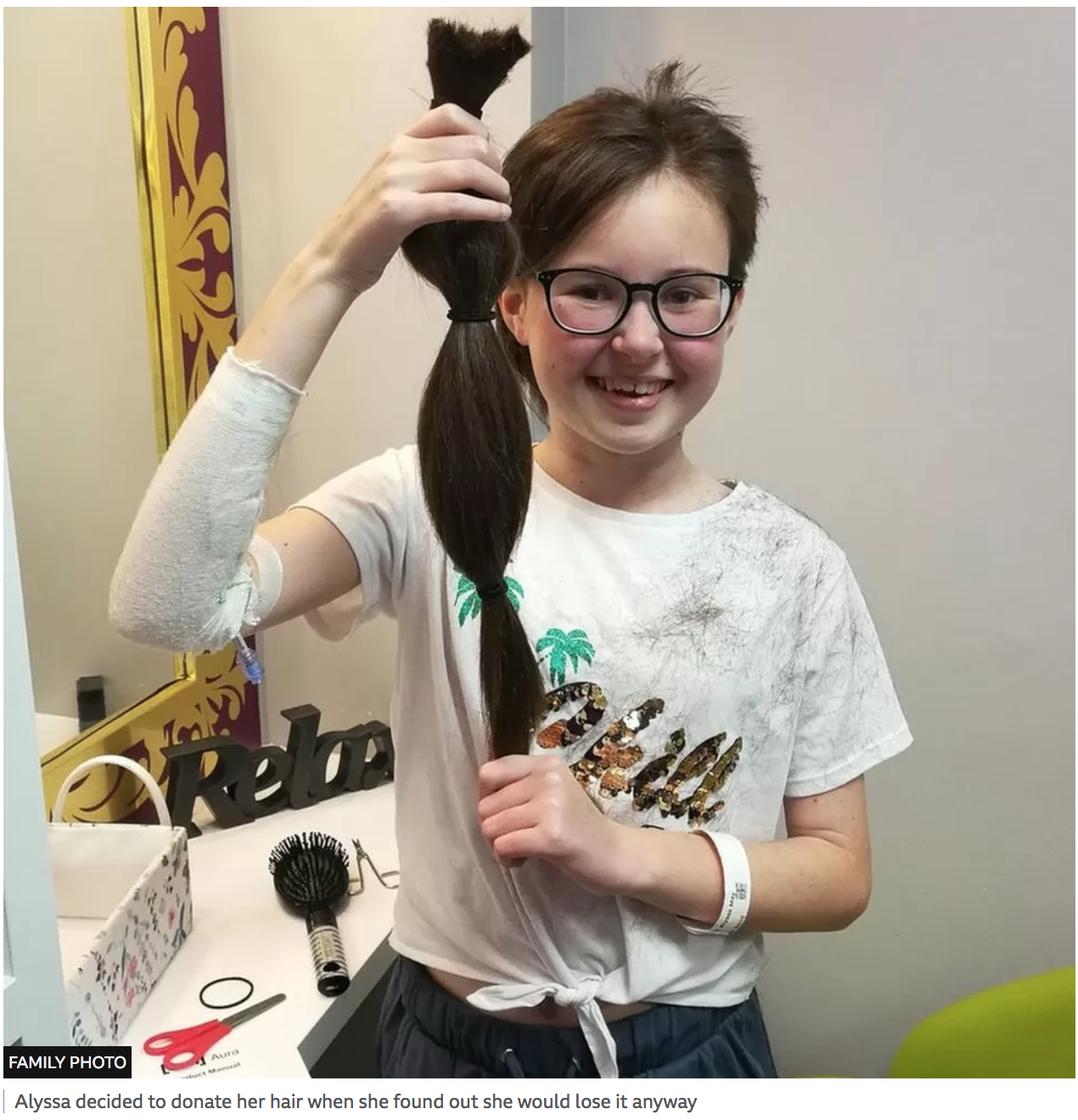
Her cancer was aggressive. Chemotherapy, and then a bone-marrow transplant, were unable to rid it from her body.
Without the experimental medicine, the only option left would have been merely to make Alyssa as comfortable as possible.
“Eventually I would have passed away,” said Alyssa. Her mum, Kiona, said this time last year she had been dreading Christmas, “thinking this is our last with her”. And then she “just cried” through her daughter’s 13th birthday in January.
What happened next would have been unthinkable just a few years ago and has been made possible by incredible advances in genetics.
The team at Great Ormond Street used a technology called base editing, which was invented only six years ago.
Bases are the language of life. The four types of base – adenine (A), cytosine (C), guanine (G) and thymine (T) – are the building blocks of our genetic code. Just as letters in the alphabet spell out words that carry meaning, the billions of bases in our DNA spell out the instruction manual for our body.
Base editing allows scientists to zoom to a precise part of the genetic code and then alter the molecular structure of just one base, converting it into another and changing the genetic instructions.
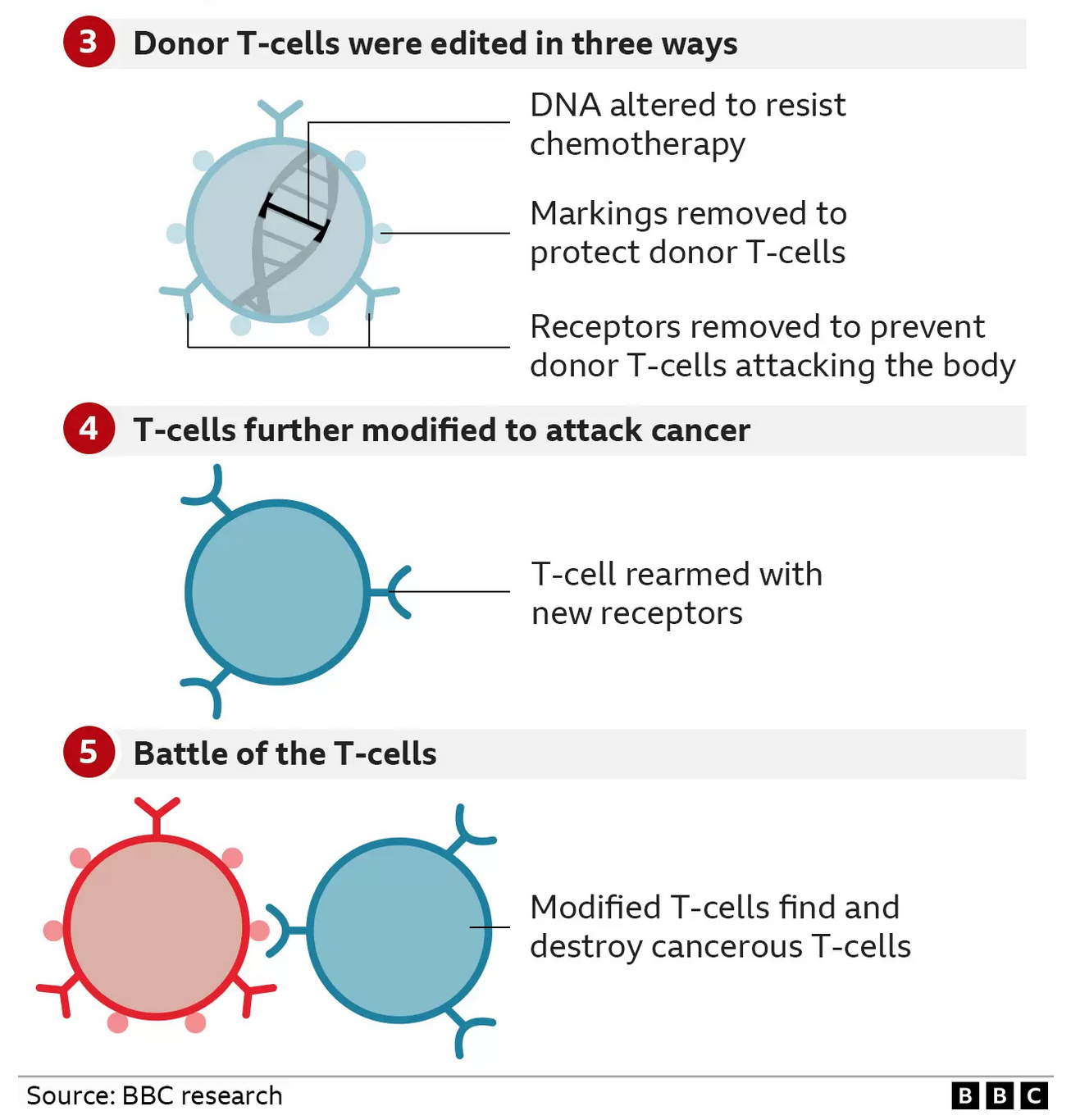
The large team of doctors and scientists used this tool to engineer a new type of T-cell that was capable of hunting down and killing Alyssa’s cancerous T-cells.
They started with healthy T-cells that came from a donor and set about modifying them.
* The first base edit disabled the T-cells targeting mechanism so they would not assault Alyssa’s body
* The second removed a chemical marking, called CD7, which is on all T-cells
* The third edit was an invisibility cloak that prevented the cells being killed by a chemotherapy drug
The final stage of genetic modification instructed the T-cells to go hunting for anything with the CD7 marking on it so that it would destroy every T-cell in her body – including the cancerous ones. That’s why this marking has to be removed from the therapy – otherwise it would just destroy itself.
If the therapy works, Alyssa’s immune system – including T-cells – will be rebuilt with the second bone-marrow transplant.
When the idea was explained to the family, mum Kiona was left thinking: “You can do that?” It was Alyssa’s decision to be the first to take the experimental therapy – which contained millions of the modified cells – in May this year.
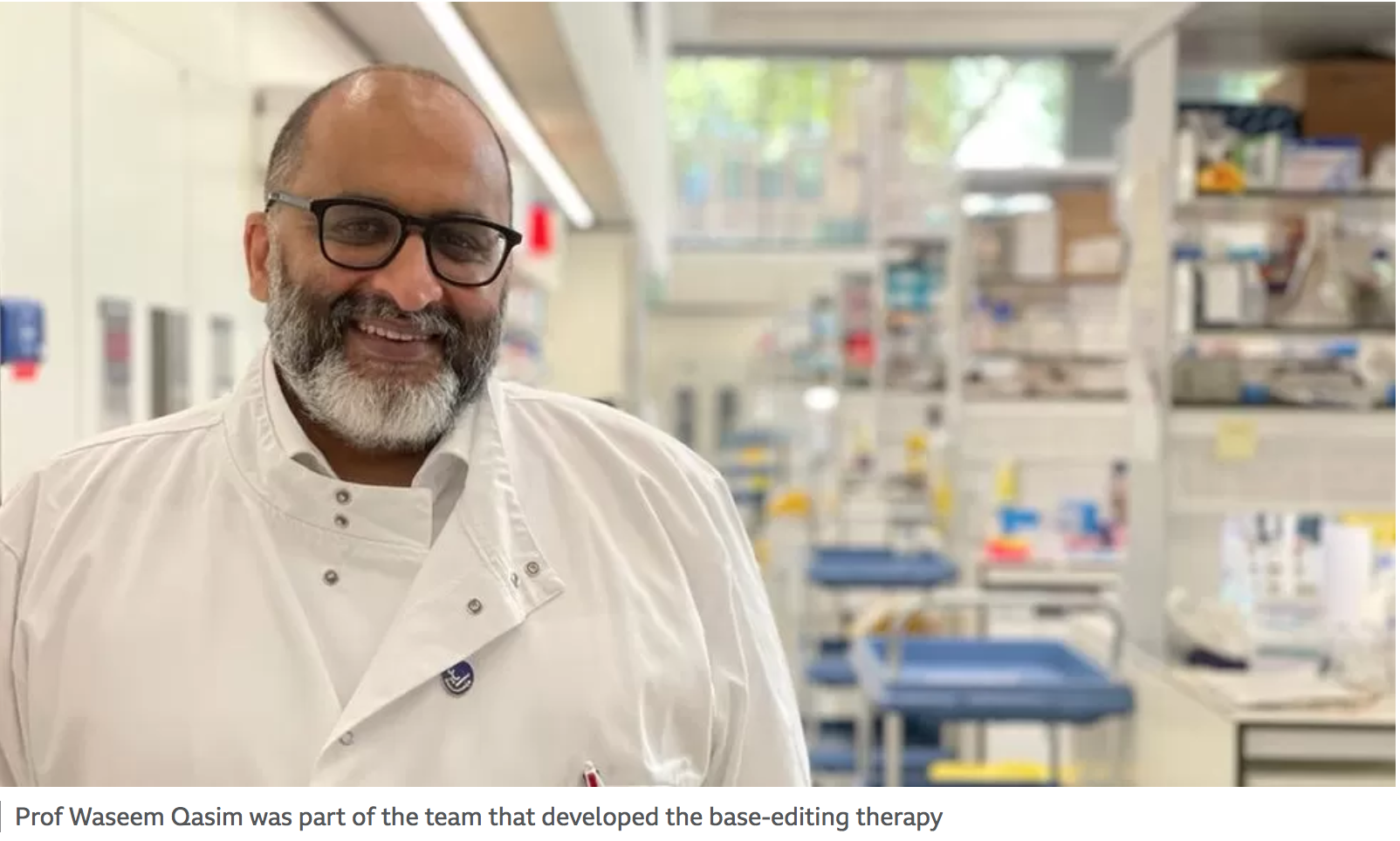
“She’s the first patient to be treated with this technology,” said Prof Waseem Qasim, from UCL and Great Ormond Street.
He said this genetic manipulation was a “very fast-moving area of science” with “enormous potential” across a range of diseases.
Alyssa was left vulnerable to infection, as the designer cells attacked both the cancerous T-cells in her body and those that protect her from disease.
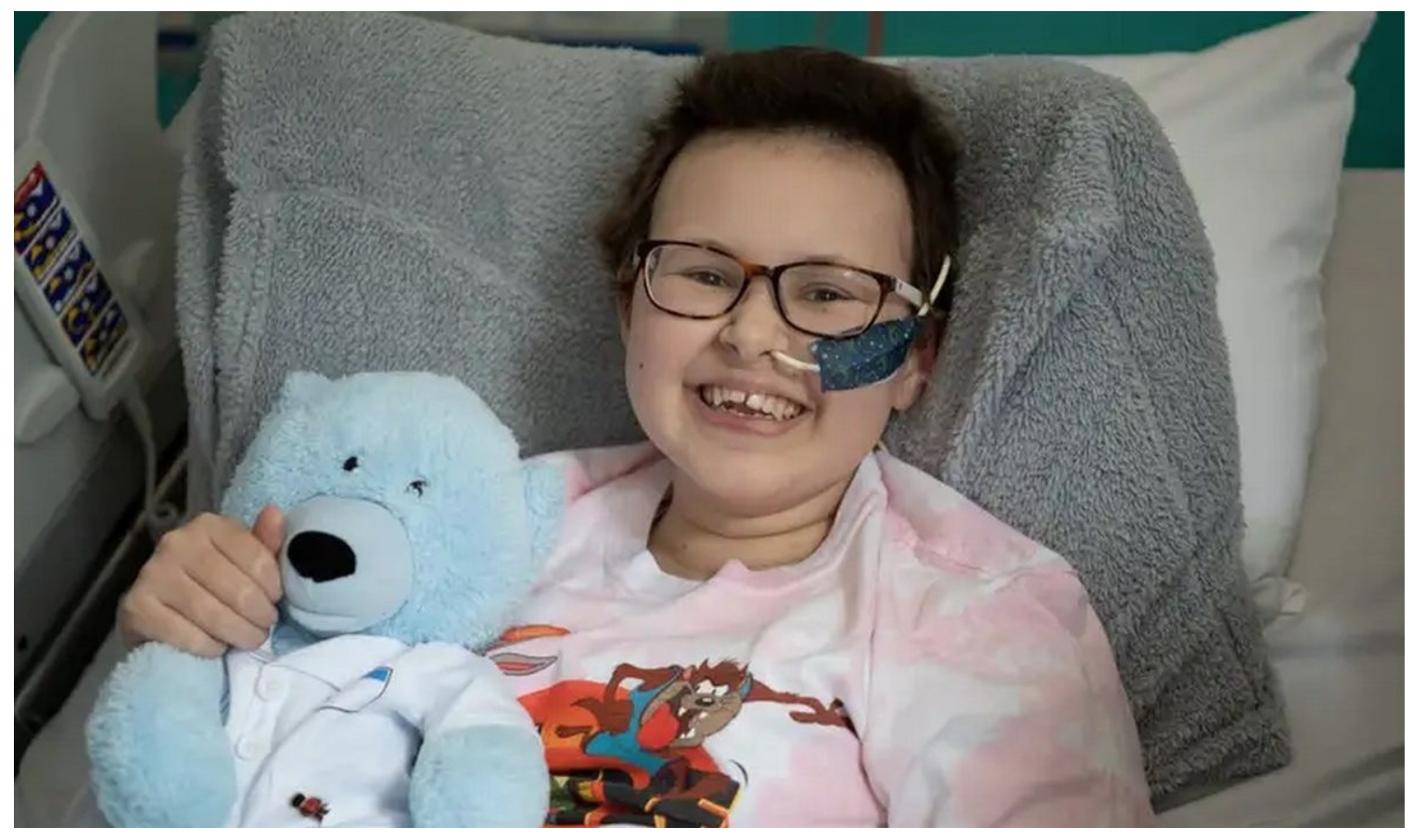
After a month, Alyssa was in remission and was given a second bone-marrow transplant to regrow her immune system.
Alyssa spent 16 weeks in hospital and couldn’t see her brother, who was still going to school, in case he brought germs in.
There were worries after the three-month check-up found signs of the cancer again. But her two most recent investigations have been clear.
“You just learn to appreciate every little thing. I’m just so grateful that I’m here now,” said Alyssa.
“It’s crazy. It’s just amazing I’ve been able to have this opportunity, I’m very thankful for it and it’s going to help other children, as well, in the future.”
She’s eyeing-up Christmas, being a bridesmaid at her auntie’s wedding, getting back on her bike, going back to school and “just doing normal people stuff”.
The family hope the cancer will never return, but are already grateful for the time it has bought them.
“To have this extra year, this last three months when she’s been home, has been a gift in itself,” said Kiona.
Dad James said: “I find it quite hard to talk about how proud we are. When you see what she’s gone through and her vitality of life she’s brought to every situation, it’s outstanding.”
Most children with a leukaemia respond to the main treatments, but it is thought that up to a dozen a year could benefit from this therapy.
Alyssa is just the first of 10 people to be given the drug as part of a clinical trial.
Dr Robert Chiesa, from the bone-marrow transplant department at Great Ormond Street Hospital, said: “It is extremely exciting. Obviously, this is a new field in medicine and it’s fascinating that we can redirect the immune system to fight cancer.”
The technology, though, only scratches the surface of what base editing could achieve.
Dr David Liu, one of the inventors of base editing at the Broad Institute, told me it was “a bit surreal” that people were being treated just six years after the technology was invented.
In Alyssa’s therapy, each of the base edits involved breaking a section of genetic code so it no longer worked. But there are more nuanced applications where instead of switching an instruction off you can fix a defective one. Sickle-cell anaemia, for example, is caused by just one base change that could be corrected.
So there are already trials of base editing under way in sickle-cell disease, as well as high cholesterol that runs in families and the blood disorder beta-thalassemia.
Dr Liu said the “therapeutic applications of base editing are just beginning” and it was “humbling to be part of this era of therapeutic human gene editing”, as science was now taking “key steps towards taking control of our genomes”.
Updated: 12-20-2022
Aggressive Leukemia Disappears In 13-Year-Old Girl Who Was First To Receive New CRISPR Treatment
In the latest CRISPR success story, a 13-year-old girl whose leukemia had not responded to other treatments now has no detectable cancer cells.
She received a dose of immune cells that were genetically edited to attack the leukemia, a method that’s been used with other cancers.
A form of cancer in the bone marrow tissue, leukemia is caused by mutated immune cells and is normally treated by killing all bone marrow cells in the patient’s body before receiving a transplant from a donor. If this falls, the Nobel Prize-winning CAR-T cell therapy can be used instead.
This was the case of Alyssa, a 13-year-old girl from the UK, who received a dose of common immune system weapons called T cells that had been modified to attack cancerous cells in her body. To avoid the extreme costs associated with this, the Great Ormond Street Hospital team at University College London further modified the donor T cells.
“This is quite remarkable, although it is still a preliminary result, which needs to be monitored and confirmed over the next few months,” said Robert Chiesa, one of the doctors treating Alyssa, in a statement released by Great Ormond Street Hospital.
While she has no detectable cancer now, it will take several years to determine whether she’s truly cancer-free.
This procedure was used before to save the life of a 1-year-old girl, Layla, also in the UK last year, and is now approved by the NHS as a treatment for people with leukemia arising from mutated B cells, another group of immune cells that can lead to the cancer.
New Scientist explains that lead researcher Waseem Qasim at University College London had applied four edits to the cells’ DNA, which puts them at risk for dangerous mutations. To circumvent this, he used a different method of the CRISPR gene editing protein that ” changes one DNA letter to another,” a technique called base-editing.
Alyssa is the first person ever to be treated with base-editing.
Updated: 6-28-2023
Researchers Discover CRISPR 2.0 Which Can Edit DNA More Precisely Than CRISPR/Cas Systems
The first RNA-guided DNA-cutting enzyme found in eukaryotes, Fanzor could one day be harnessed to edit DNA more precisely than CRISPR/Cas systems.
A team of researchers led by Feng Zhang at the McGovern Institute for Brain Research at MIT and the Broad Institute of MIT and Harvard has uncovered the first programmable RNA-guided system in eukaryotes — organisms that include fungi, plants, and animals.
Related:
In a study published today in Nature, the team describes how the system is based on a protein called Fanzor. They showed that Fanzor proteins use RNA as a guide to target DNA precisely, and that Fanzors can be reprogrammed to edit the genome of human cells.
The compact Fanzor systems have the potential to be more easily delivered to cells and tissues as therapeutics than CRISPR-Cas systems, and further refinements to improve their targeting efficiency could make them a valuable new technology for human genome editing.
CRISPR-Cas was first discovered in prokaryotes (bacteria and other single-cell organisms that lack nuclei) and scientists including those in Zhang’s lab have long wondered whether similar systems exist in eukaryotes. The new study demonstrates that RNA-guided DNA-cutting mechanisms are present across all kingdoms of life.
“CRISPR-based systems are widely used and powerful because they can be easily reprogrammed to target different sites in the genome,” says Zhang, senior author on the study, the James and Patricia Poitras Professor of Neuroscience in the MIT departments of Biological Engineering and Brain and Cognitive Sciences, an investigator at MIT’s McGovern Institute, a core institute member at the Broad Institute, and a Howard Hughes Medical Institute investigator. “This new system is another way to make precise changes in human cells, complementing the genome editing tools we already have.”
Searching The Domains Of Life
A major aim of the Zhang lab is to develop genetic medicines using systems that can modulate human cells by targeting specific genes and processes. “A number of years ago, we started to ask, ‘What is there beyond CRISPR, and are there other RNA-programmable systems out there in nature?’” says Zhang.
Two years ago, Zhang lab members discovered a class of RNA-programmable systems in prokaryotes called OMEGAs, which are often linked with transposable elements, or “jumping genes,” in bacterial genomes and likely gave rise to CRISPR-Cas systems.
That work also highlighted similarities between prokaryotic OMEGA systems and Fanzor proteins in eukaryotes, suggesting that the Fanzor enzymes might also use an RNA-guided mechanism to target and cut DNA.
In the new study, the researchers continued their work on RNA-guided systems by isolating Fanzors from fungi, algae, and amoeba species, in addition to a clam known as the northern quahog.
Co-first author Makoto Saito of the Zhang lab led the biochemical characterization of the Fanzor proteins, showing that they are DNA-cutting endonuclease enzymes that use nearby non-coding RNAs known as ωRNAs to target particular sites in the genome. It is the first time this mechanism has been found in eukaryotes, such as animals.
Unlike CRISPR proteins, Fanzor enzymes are encoded in the eukaryotic genome within transposable elements, and the team’s phylogenetic analysis suggests that the Fanzor genes have migrated from bacteria to eukaryotes through so-called horizontal gene transfer.
“These OMEGA systems are more ancestral to CRISPR and they are among the most abundant proteins on the planet, so it makes sense that they have been able to hop back and forth between prokaryotes and eukaryotes,” says Saito.
No Collateral Damage
To explore Fanzor’s potential as a genome editing tool, the researchers demonstrated that it can generate insertions and deletions at targeted genome sites within human cells.
The researchers found the Fanzor system to initially be less efficient at snipping DNA than CRISPR-Cas systems, but by systematic engineering, they introduced a combination of mutations into the protein that increased its activity 10-fold.
Additionally, unlike some CRISPR systems and the OMEGA protein TnpB, the team found that a fungal-derived Fanzor protein did not exhibit “collateral activity,” where an RNA-guided enzyme cleaves its DNA target as well as degrading nearby DNA or RNA. The results suggest that Fanzors could potentially be developed as efficient genome editors.
Co-first author Peiyu Xu led an effort to analyze the molecular structure of the Fanzor/ωRNA complex and illustrate how it latches onto DNA to cut it.
Fanzor shares structural similarities with its prokaryotic counterpart CRISPR-Cas12 protein, but the interaction between the ωRNA and the catalytic domains of Fanzor is more extensive, suggesting that the ωRNA might play a role in the catalytic reactions.
“We are excited about these structural insights for helping us further engineer and optimize Fanzor for improved efficiency and precision as a genome editor,” said Xu.
Like CRISPR-based systems, the Fanzor system can be easily reprogrammed to target specific genome sites, and Zhang said it could one day be developed into a powerful new genome editing technology for research and therapeutic applications.
The abundance of RNA-guided endonucleases like Fanzors further expands the number of OMEGA systems known across kingdoms of life and suggests that there are more yet to be found.
“Nature is amazing. There’s so much diversity,” says Zhang. “There are probably more RNA-programmable systems out there, and we’re continuing to explore and will hopefully discover more.”
The paper’s other authors include Guilhem Faure, Samantha Maguire, Soumya Kannan, Han Altae-Tran, Sam Vo, AnAn Desimone, and Rhiannon Macrae.
Support for this work was provided by the Howard Hughes Medical Institute; Poitras Center for Psychiatric Disorders Research at MIT; K. Lisa Yang and Hock E. Tan Molecular Therapeutics Center at MIT; Broad Institute Programmable Therapeutics Gift Donors; The Pershing Square Foundation, William Ackman, and Neri Oxman; James and Patricia Poitras; BT Charitable Foundation; Asness Family Foundation; Kenneth C. Griffin; the Phillips family; David Cheng; Robert Metcalfe; and Hugo Shong.
Related Article:
Why It’s So Hard To Find Dumbbells In The US
History of Alchemy From Ancient Egypt To Modern Times
Gyms Reopening May Not Facilitate Coronavirus Infections, Study Finds
These Are Your Rights When A Barber, Gym Owner Or President Trump Asks You To Sign A COVID-19 Waiver
As Coronavirus Lockdowns Lift, How Ready Are Gyms To Reopen?
Leery of Joining A Health Club? Today’s Gyms Are Good For Physical, And Fiscal, Fitness
Does Getting Stoned Help You Get Toned? Gym Rats Embrace Marijuana
Spy Gadgets For Grow Facilities / Weed Dispensaries (#GotBitcoin?)








Leave a Reply
You must be logged in to post a comment.